 
And indeed, the era of wheat genomics has begun. Following this publication and the emergence of new sequencing technologies, the sequencing and assembly of all other known wheat relatives became much easier to handle. This was a breakthrough in deciphering the large and complex genome of bread wheat and its relatives. In 2017, first genome draft of wild emmer wheat was published ( Avni et al., 2017), after a joint effort between industry and academic research groups that has led to the development of an assembly algorithm capable of dealing with highly repetitive DNA sequences. 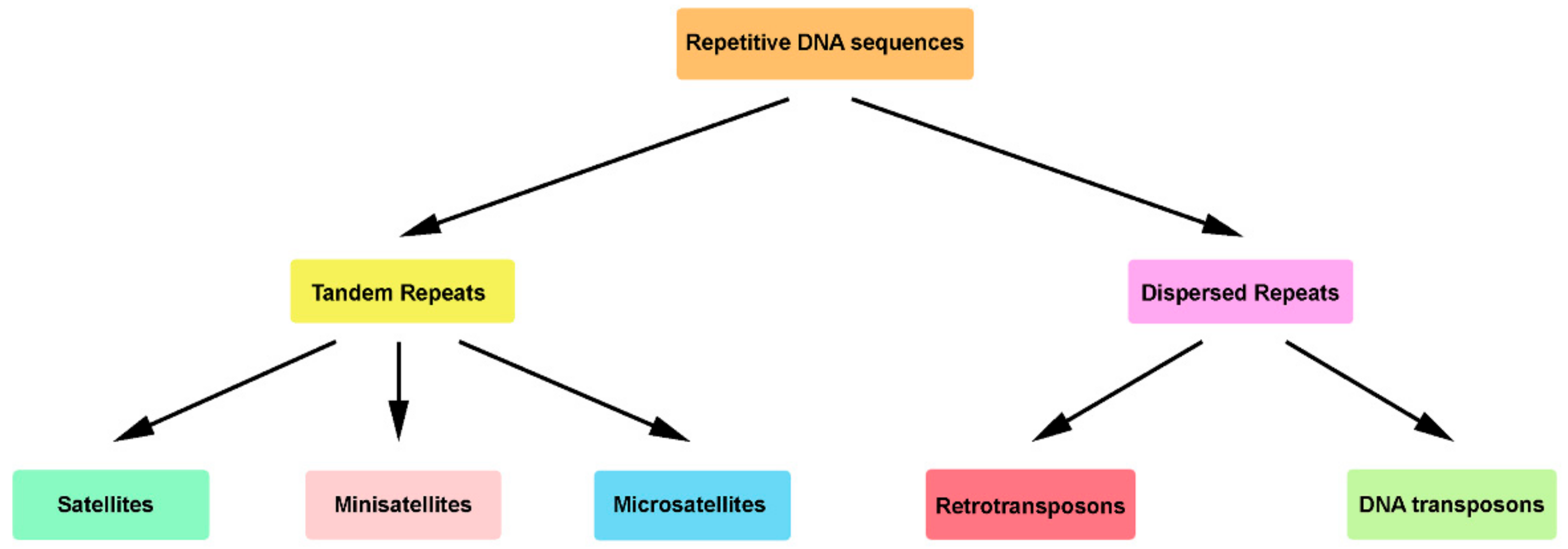
This genome draft gave the wheat community a first real glance into the complicated wheat genome. ( Mayer et al., 2014) and was based on chromosome-based sequencing. The first genome draft of bread wheat was published by Mayer et al. The Era of Wheat Genomicsĭuring the last decade, rapid improvement of DNA sequencing and assembly methods enabled the generation of whole genome assemblies for bread wheat and some of its progenitors. The repetitive nature of the wheat genome, together with its huge size – 17 Gbp, have delayed the generation of the bread wheat genome draft when many other organisms (both animals and plants) have already been sequenced ( Bolger et al., 2014 Uauy, 2017 Guan et al., 2020). Each class of TEs can be further divided into subclass, orders, super-families, families, and subfamilies, and includes both autonomous and non-autonomous TEs. TEs are DNA fragments capable of increasing their copy number within the host genome mostly through copy and paste (Class I, retrotransposons) or cut and paste (Class II) mechanisms ( Wicker et al., 2007). The repetitive nature of the wheat genome is mainly the result of a high transposable elements (TEs) content. The major obstacle in creating a reference genome draft for bread wheat was the assembly of contigs made from mostly (over 80%) repetitive sequences. This allopolyploidization event resulted in the speciation of bread wheat ( Triticum aestivum), which today is among the world’s most widely grown crops, providing nearly 20% of daily human caloric intake ( Avni et al., 2017 Appels et al., 2018 Pont et al., 2019).Īlthough wheat is a highly important crop, the challenges in wheat genomics have led to a relatively slow advancement in this field during the beginning of the next generation sequencing (NGS) era as reviewed by Guan et al. 
The second allopolyploidization event that occurred ~10,000 year ago was the result of hybridization between a tetraploid from the durum wheat lineage and the D genome donor ( Aegilops tauschii). The initial domestication and cultivation of wild emmer gave rise to the tetraploid lineage, that following selection resulted in the free-threshing durum wheat, T. About 0.5 MYA, a hybridization of the A and the B genome donors that was followed by polyploidization, led to the speciation of the allotetraploid wild emmer, T. This raises fundamental questions with regards to the differences and commonalities between PcG- and DNA methylation-mediated silencing and the role and mechanistic determinants of PcG targeting at TEs.Bread wheat ( Triticum aestivum) is a relatively young allohexaploid species, which has been generated by two subsequent allopolyploidization events that followed the divergence of three diploid wild ancestors: Triticum urartu (donor of the A genome), a species from section Sitopsis, a relative of today’s Aegilops speltoides (donor of the B genome) and Aegilops tauschii (donor of the D genome) ( Feldman and Levy, 2012 Pont et al., 2019). In particular, many TE sequences, upon their loss of DNA methylation and/or in certain cell-types, display H3K27me3 marks and even, more rarely, in the presence of DNA methylation. Nevertheless, there is a growing body of evidence for a functional and mechanistic interplay between these pathways. Thus, PcG and DNA methylation have been generally seen as mutually exclusive, specialized systems for the transcriptional silencing of genes and TEs respectively, in plants and mammals. 
Inversely, in most model organisms, the highly conserved Polycomb Group (PcG) proteins, namely Polycomb Repressive Complex 2 (PRC2), which deposits histone H3 Lysine 27 trimethylation (H3K27me3), are a hallmark of transcriptional repression associated with protein- and miRNA-coding genes involved, in particular, in development. DNA methylation negatively controls TE expression in a stable manner and typically is not involved in silencing protein coding genes. Epigenetic regulation of Transposable Elements (TE)ĭNA methylation (5-methylcytosine) is a hallmark of TEs in many organisms, including plants in which it is established by RNA silencing pathways (RNA-directed DNA methylation) and associated with histone H3K9 dimethylation. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
